Tuhina Banergee
This year, like every other year, one in six Americans will get sick from a foodborne illness. That’s roughly 48 million people. According to Centers for Disease Control (CDC), 128,000 of those people will be hospitalized and 3,000 of them will die.
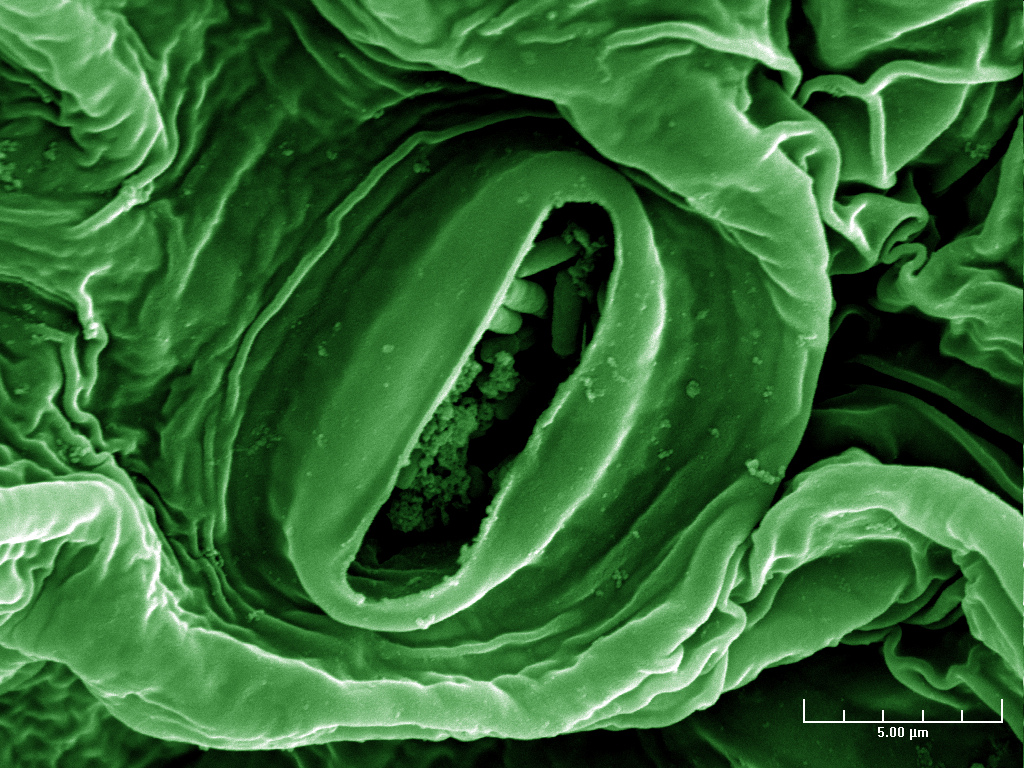
E. coli is a large group of bacteria that’s commonly found in the intestines of humans and animals. Most strains of the slender, rod-shaped bacteria are pretty harmless. They’re part of a healthy human gut microbiome. But when certain strains that normally reside in the guts of animals end up in our human intestines by way of contaminated food, they can become pathogenic and morph into an infection. That’s when we collide with the E. coli of Chipotle or Costco fame—E. coli 0157: H7—the kind of virulent strain that makes people really, really sick.
E. coli 0157: H7 is like a rampage strain, an outlaw among outbreaks. It’s been responsible for some of the worst in recent memory, including the notorious Jack in the Box outbreak of 1993, which sickened more than 500 people and killed four, at least three of them children.
In 2015, United States Department of Agriculture (USDA) conducted a study on how much economic damage the United States suffers annually as a result of foodborne illnesses. It found that, of the 48 million cases of foodborne illness diagnosed every year, a specific pathogen could be identified in only 20 percent of them. And over 90 percent of those cases are caused by just 15 pathogens, including E. coli 0157.
The total bill tallies up this way: 15 wily, microscopic pathogens are behind nearly 9 and a half million illnesses. Those illnesses cost the U.S more than $15 billion in medical expenses and lost productivity every year.
Now, back to that identification business for a moment. Obviously, it helps whenever we can identify a strain of bacteria before it makes people sick. But conventional methods of detection—agar-culturing, for instance—can take at least 24 to 48 hours to yield results, and don’t always catch small traces of E. coli, particularly 0157: H7.
In recent years, PCR (polymerase chain reaction), a test based on the process of DNA replication and conducted in vitro, became a relatively cost-effective way to get same-day results on bacterial identification. But PCR requires lab use and trained technicians, and is highly sensitive to even the slightest procedural mistake, which can produce erroneous results. This makes detecting large-scale outbreaks—in food manufacturing, for example—a less-than-ideal use for PCR.
But what if detecting a foodborne pathogen was as simple as, say, scanning a food label for GMOs in the grocery store aisle? And what if getting an accurate result was just as fast?
Well, we might be a self-driving car ride away from that reality. Scientists at the Kansas Polymer Research Center at Pittsburg State University have developed a new nanosensor that can rapidly detect the presence of E. coli in food or water (is an hour fast enough for you?) and it may pave the way for prevention of outbreaks—not just of E. coli, but a range of other foodborne illnesses, too.
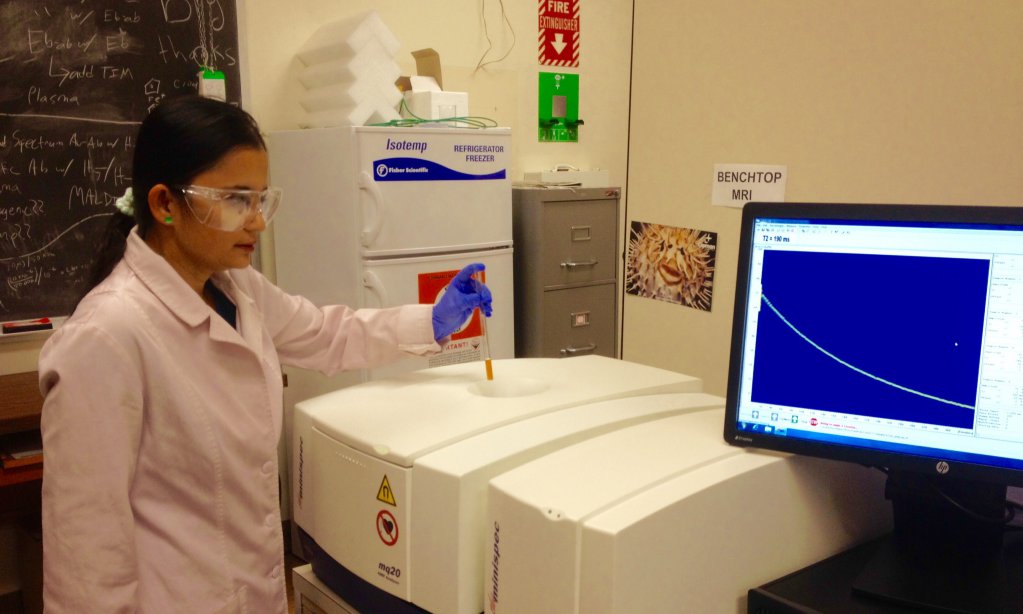
The study, which was jointly funded by K-INBRE, the Kansas Soybean Commission, and PSU’s Polymer Chemistry Startup Fund, was recently published in the journal ACS Infectious Diseases. Tuhina Banerjee, one of the scientists working on the project, says there are two existing rapid detection methods—magnetic resonance and fluorescence—that can detect minute and mass quantities of bacteria, respectively. Banerjee and her colleagues wondered whether combining these methods would create a better, more dynamic sensor. It worked.
“When we coupled both these modalities together, our nanosensors could detect from very, very low range, to very high range [of bacteria]. That’s the whole beauty of this sensor,” Banerjee says. In other words, a single sensor could be useful in detecting both trace amounts of E. coli (which current methods struggle to do) and mass amounts, like in a body of water, making it as useful in a large-scale U.S. meat industry outbreak as it would be in a waterborne illness crisis in India.

Fluorescence is capable of detecting much larger quantities of bacteria, while magnetic resonance detects smaller amounts
Banerjee says she was inspired to use E. coli 157: H7 because she’s got specific connections to that particular pathogen. “Right now I’m in Kansas and actually, I came from India. So I really wanted to do something,” she says. “In India there’s cholera and so many kinds of bacterial infections are really prevalent. And in Kansas, you have lots of cases where there are a lot of cattle and the feces of cattle are crossed with drinking water.”
But this new nanosensor wasn’t developed solely for use in detecting E. coli. Banerjee says it’s more versatile than that. “Right now the problem is Zika. We can use the conjugation chemistry we have used for this particular case and we can tweak it for use in other applications.”
Of course, the chemistry itself is only mortar. What we need next are bricks. That’s where engineers come in. “Right now what we have to rely on, because it’s just the basic steps, is the fluorescent plate reader and the benchtop MRI machine,” Banerjee says. “These two instruments are required” to read the sensor results. But she envisions working with engineers to implement this nanosensor technology in the form of a chip for use in a phone or nanodevice.
“So suppose you have that bacteria in beef. You extract a sample, you take a few microliters of the solution, and you use your nanosensor to scan it and if it shows a change, that means you have the infection there and something needs to be done,” says Banerjee.
If successful, the new method could be a milestone in the fight against foodborne illness: the ability to detect the presence of bacteria on-site, in an hour, even before it hits shelves and long before anyone gets sick.